Optimization of the sample preparation method for cell-free DNA methylome analysis
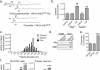
To optimize the sample preparation method for cfDNA methylome analysis, we first examined the impact of anticoagulant type on the cfDNA yields. For this purpose, whole blood was collected into EDTA (n = 3) and CPT (n = 4) tubes. Plasma was fractionated and cfDNAs were isolated from each sample by using 1,000 μl volume of plasma. The cfDNA yields varied between 2.05 and 17.08 ng. No statistically significant differences were detected in the cfDNA yields between EDTA or CPT plasma (paired t-test p-value 0.20, Supplementary Fig. S1). We then examined impact of plasma volume on the cfDNA yields. For this purpose, cfDNAs were isolated from 1,000, 750 and 500 μl of EDTA plasma with Qiagen QIAamp Circulating Nucleic Acid kit that is optimized for up to 5 ml plasma volumes. When using 500 μl of…